|
Registration fee includes coffee breaks, lunches, and course materials. Late registration participants will be accepted upon availability. Upon acceptance to the course, early registration participants have until the 15th June to complete the payment, otherwise waiting list participants will be invited to the course. Registration is conditional, course acceptance depends on motivation and date of registration, and will be informed by 3rd June. The course modules are independent of each other, therefore you are free to choose the module(s) of your preference.Įach module requires a minimum number of 10 participants and the maximum number of participants is 25. Knowledge about the specific image analysis software to be used in the course is not required, however, some prior knowledge about image analysis is beneficial. Therefore, participants are free to choose the image analysis tools more convenient to their own work.Īndré Maia and María Gómez | on BioImage Analysis for High Content ScreeningĪndré Maia and María Gómez | participant may choose the following modules: Module Experimental data will be used as datasets for image segmentation and quantification. Fundamentally practical, in this modular course, participants will get acquainted with several open-source image analysis software (ImageJ/Fiji, ilastik, CellProfiler, CellProfiler Analyst, Cytoscape and IDEAS®) designed to deal with large amounts of images. 
Images contain diverse and valuable quantifiable data, however, many researchers lack the knowledge to extract those data in a practical, fast and repetitive way. High Content Screening generates massive amounts of image-based data in the biosciences field. I3S - Instituto de Investigação e Inovação em Saúde da Universidade do Porto In light of the current COVID-19 crisis, this course has been cancelled. Please note that if you are not eligible for a University of Cambridge Raven account you will need to book or register your interest by linking here.Workshop on BioImage Analysis for High Content Screening The training room is located on the first floor and there is currently no wheelchair or level access available to this level. We will also briefly discuss the basic principles of supervised machine learning with CellProfiler Analyst in order to score complex and subtle phenotypes. We will show how CellProfiler can be used to analyse a variety of types of imaging experiments. It is selected and loaded upon startup of CPA. This file can be stored anywhere on your computer. The properties file is a plain text file that contains the configuration information necessary for CPA to access your data and images. This course will introduce users to the free open-source image analysis program CellProfiler and its companion data exploration program CellProfiler Analyst. Setting up the properties file CellProfiler Analyst 2.2.1 documentation. From small-scale microscopy experiments to time-lapse movies and high-throughput screens, automatic image analysis is more objective and quantitative and less tedious than visual inspection. Microscopy experiments have proven to be a powerful means of generating information-rich data for biological applications. University Information Services - Staff Learning & Development.University Information Services - Digital Literacy Skills.Social Sciences Research Methods Programme.Schools of Physical Sciences & Technology.Schools of Arts, Humanities & Social Sciences.PPD Personal and Professional Development. 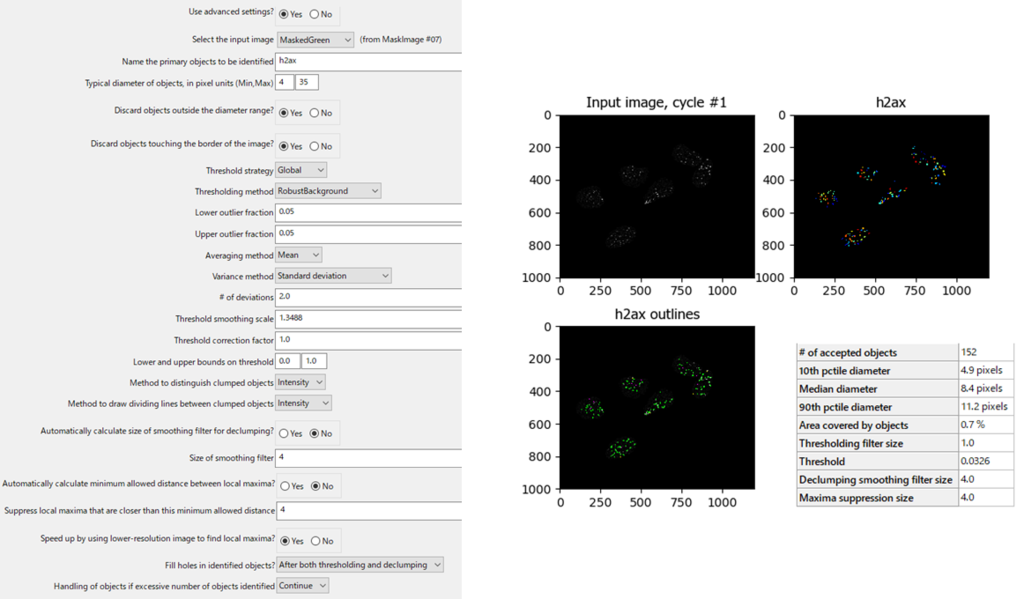
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
